Today we met to discuss the assemblies we have got so far using ABySS-2.0.2-k128 as an assembler and running three different k-mer sizes; 31, 48 and 55. These are the statistics for each:
k-mer of 31 (Run by Panos)
n n:500 L50 min N80 N50 N20 E-size max sum name
3982947 28472 7791 500 710 1225 2258 1585 9621 31.15e6 3pandina_assembled-unitigs.fa
k-mer of 48 (Run by me)
n n:500 L50 min N80 N50 N20 E-size max sum name
2840597 43385 13871 500 616 897 1525 1133 5973 38.34e6 p-andina-assembly-untrimmed-unitigs.fa
k-mer of 55 (Run by Maria)
n n:500 L50 min N80 N50 N20 E-size max sum name
3321446 42326 16004 500 569 718 1013 818 3689 31.01e6 Assembly7_k15-unitigs.fa
To proceed, each of us will run the assembly on different k-mer sizes as 31:4:90. Panos will assemble with k-mer sizes 31, 35, 39, 43, and 47, I will assembly using k-mer sizes 51, 55, 59, 63, and 67, and Maria will assemble with k-mer sizes of 71, 75, 79, 83, and 87. This will be done un the untrimmed data. The goal is to then plot different k-mer sizes against different evaluation parameters to assess the effect of k-mer size on assembly quality.
The contig-file based on the k-mer size that resulted in highest quality will be used further in the project (for example if we need to use a scaffolder with contigs as input).
I will discuss the different parameters in another post when we have tested with different k-mer sizes.
What bothers me a lot is that we are not obtaining the same files as other groups are, even though we run the same version of ABySS and basically same commands, except for the fact that we only have single-reads, and other groups have paired end reads. This leaves me thinking that the problem is some of the following:
- ABySS thinks that the contig results are too bad to continue with the scaffolding
- Scaffolding is only performed by ABySS if we specify the different paired end reads (and not just single reads, i.e. the
-seoption is not working properly).
This means we have to consider using a scaffolder which we first did not think was needed when seeing that other groups obtain .scaffold-files after running ABySS. However, when reading in different forums and documentation pages, it seems like scaffolding is not possible with only single reads as the scaffolder needs the paired ends to connect the contigs into an actual scaffold. During the meeting, we decided to e-mail Lars Arvestad and ask for tips on how to proceed with the assembly.
We also decided to divide the work a bit since we are stuck on the assembly-part of the project. After running the assembly with different k-mer sizes, I will continue alone with the assembly and run all k-mers again but in the trimmed data, and potentially do the scaffolding-part (depending on what answers we get from Lars), and in parallel, Maria and Panos will start working with the CHS-genes and BLAST, just to make sure we get SOME results even though we are stuck on the assembly part.
Tonight, I will run ABySS on the k-mers as stated above. One problem though is that each run generates a lot of data, and this takes up quite a lot of memory. The initial plan was to make a for-loop that creates a new folder for each k-mer, with the output file subsequently ending up in the corresponding folder). However, as we only have 36 Gb, I will have to remove the output files after each run and after extracting the stats (which is the only thing that is interesting for now), and then start a new run with a new k-mer. When we have finally decided which k-mer best suits our assembly, we can easily generate that data again. This is just for testing.
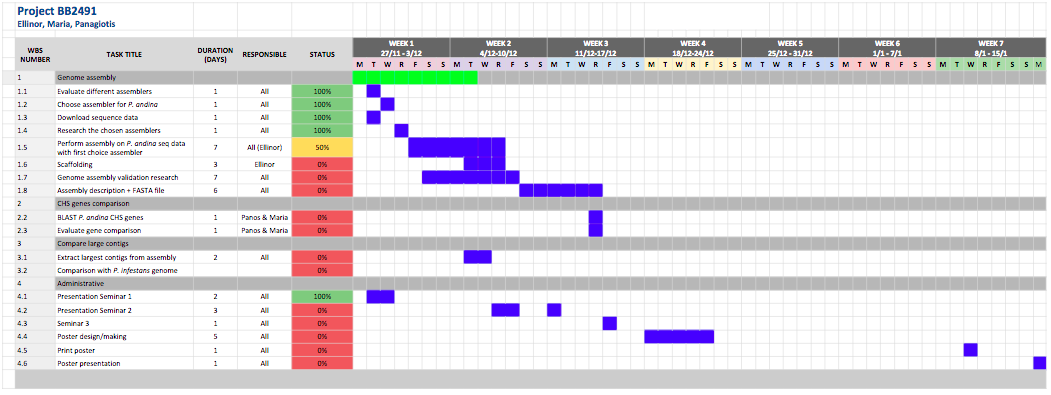
Because if the issues we have had with the assembly (which we first planned 3-4 days for) our project plan has changed a bit. Below is the current, and most updated Gantt schedule:
I will update tomorrow with the results from the assembly runs, and hopefully after reading an answer from Lars which could help us to proceed with the assembly part.